Recently,a research team led by ProfessorYang Dayongfrom the School of Chemical Engineering and Technology at Tianjin University successfully developed a DNA hybridization chain reaction (HCR)-based strategy that can achieve spatiotemporally programmable assembly of DNA under nanoconfinement for precise siRNA delivery. The study was published in Nature Communications(http://www.tju.edu.cn/info/1026/4003.htm).
In living cells, the physicochemical characteristics of confinement essentially play pivotal roles, such as stabilizing the fold conformation of biomolecules, enhancing biochemical reactivity, affecting biochemical equilibrium, and achieving simultaneous occurrence of multi-bioprocesses in confined spaces.
In recent years, DNA has been used as a versatile building block to assemble functional nanostructures for a wide range of applications due to the unparalleled sequence programmability, molecular recognition, and biological functions. In particular, DNA nanostructures have been proven as promising nanocarriers for delivery of nucleic acid drugs. Whereas, DNA assembly under polymeric nanoconfinement has not been explored for the delivery of nucleic acid drugs till now.
The system newly developed by Yang’s research group achieves spatiotemporally programmable assembly of DNA under nanoconfinement for precise siRNA delivery. The polymeric nanoframework with DNA as cross-linker is prepared via precipitation polymerization. By using the potential energy stored in the loops of DNA hairpins to overcome the steric effect under nanoconfinement, the cross-linker DNA in the polymeric nanoframework is designed to initiate the cascade hybridization of two DNA hairpins, thus achieving efficient loading of siRNA in the polymeric nanoframework. Moreover, functional groups such as tumor-targeting phenylboronate are easily decorated on the polymeric nanoframework, which can overcome complexity–scalability–error issues of DNA-only nanosystems, improving the in vivo gene delivery efficiency. By virtue of these features, siRNA loading DPNF with good physiological stability, enhanced cellular uptake, and controlled siRNA release property were obtained; consequently, efficient gene knockdown was achieved both in vitro and in vivo.
Given the abundance of DNA-templated reactions and versatile polymer systems, the spatiotemporally programmable assembly of other gene drugs under nanoconfinement is easily achieved, which provides extensible strategy to engineer smart nucleic acid nanoplatform for precision medicine.
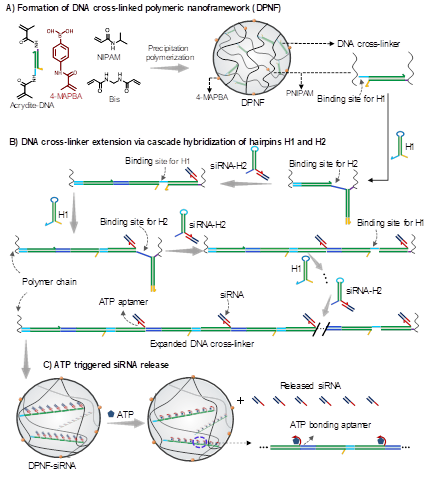
Molecular design.Schematic diagram of(A)the preparation of DPNF,(B) DNA cross-linker extension for siRNA loading via cascade hybridization of DNA hairpins in DPNFs, and (C) ATP triggered release of siRNA from the polymeric nanoframework.
By the School of Chemical Engineering and Technology
Editor: Eva Yin